Previous issue | Next issue | Archive
Volume 10 (2); March 25, 2020![]() [Booklet]
[Booklet]
 Research Paper
Research Paper
Comparative analysis of the videothoracoscopic interventions results
| Comparative analysis of the videothoracoscopic interventions results |
Eshonkhodjaev OD, Ibadov RA, Bobayev UN and Ismailov BA.
J. Life Sci. Biomed., 10(2): 10-16, 2020; pii:S225199392000002-10
DOI: https://dx.doi.org/10.29252/scil.2020.jlsb2
Abstract
Aim. This study was done to determine the feasibility and effectiveness of the proposed method of thoracoscopic hemostasis and aerostasis. Methods. The study included 85 patients operated for bullous lung disease, closed chest injury and penetrating chest wounds in the Lung and Mediastinum surgery department of the Republican Specialized Scientific and Practical Medical Center of Surgery named after Academician V.Vakhidov for the period from 2015 to 2019. Total of 33 patients made up the main group: thoracoscopy using the proposed technique and 52 patients for the comparison group: thoracoscopic aerostasis was performed using known methods. In 21 (40.4%) cases of comparison group, we performed video-assisted thoracoscopic (VATS) excision and suturing with pleurodesis; in 14 (26.9%) cases – VATS with stitching of a lung wound. VATS excision and flashing of bullae of the lung using a stapler was performed in 19.2% (10 of 52) cases of the comparison group and 24.2% in the main group, where all VATS were supplemented with Geprotsel gel application. Results. Using the Geprotsel in VATS interventions allowed to reduce the necessity of lung tissue stitching from 67.7% to 27.3%, respectively, to limit excision in 36.4% of patients, to achieve complete tightness after hardware stitching (χ2 - 17.304; Df=3; p<0.001), which generally leveled the risk of postoperative pneumonia and impaired hemostasis. Recommendation. We suggest that applying Geprotsel gel during VATS for lung tissue damages allows to reduce the application of additional sutures, improve the efficiency of minimally invasive operations with a decrease in the frequency of postoperative disorders of aero- and hemostasis.
Keywords: Lung Pathology, Video-Assisted Thoracoscopy, Geprotsel, Hemostasis, Aerostasis
[Full text-PDF] [HTML] [ePub] [XML] [Export citation as BibTeX, RIS & EndNote] [How to Cite] [Google Scholar] [Dimensions ID]
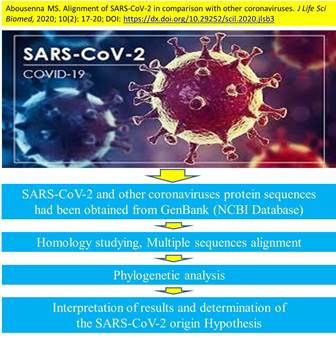
Alignment of SARS-CoV-2 in comparison with other coronaviruses
| Alignment of SARS-CoV-2 in comparison with other coronaviruses |
Abousenna MS.
J. Life Sci. Biomed., 10(2): 17-20, 2020; pii:S225199392000003-10
![]() DOI: https://dx.doi.org/10.29252/scil.2020.jlsb3
DOI: https://dx.doi.org/10.29252/scil.2020.jlsb3
Abstract
Introduction. Severe Acute Respiratory Syndrome Coronavirus 2 (SARS-CoV-2) is currently declared as pandemic according to WHO. It was initially detected in China and then rapidly transmitted to most world territories. The SARS-CoV-2 has an ambiguous origin, with unique properties, pathogenesis and transmission rate, thus making its prevention and control a difficult task. Aim. In the present study, we investigated the origin hypotheses through conducting multiple alignments and phylogenetic analysis for surface glycoprotein and complete genome of SARS-CoV-2 in comparison with other coronaviruses of different species. All the data used in this study were obtained from NCBI online database and analyzed using Blast tool. The alignment and phylogenetic analysis of SARS-CoV-2 surface glycoprotein in comparison with spike glycoprotein of Bat coronavirus RaTG13, Pangolin coronavirus, Bat SARS-like coronavirus, SARS-CoV, BCoV, IBV, ECoV, MHV-JHM, MERS-CoV, CCoV, HCoV-229E and FCOV indicated close identical matching to spike protein for Bat coronavirus RaTG13 and Pangolin coronavirus isolate MP789. The similarity was 97.41% and 96.67%, respectively. Also, multiple alignments of complete genome for SARS-CoV-2 and Bat coronavirus RaTG13 showed a significant similarity of 96.11%. Recommendation. Therefore, these relevant results strongly recommend the origin hypothesis of SARS-CoV-2 from Bat coronavirus RaTG13. The nature of evolution is considered to be natural selection.
Keywords: Severe Acute Respiratory Syndrome, SARS-CoV-2, COVID-19, Coronavirus Disease 2019, Alignment, Phylogenetic analysis.
[Full text-PDF] [HTML] [ePub] [XML] [Export citation to BibTeX, RIS & EndNote] [How to Cite] [Google Scholar] [Dimensions ID]


