Previous issue | Next issue | Archive
Volume 10 (4); July 25, 2020![]() [Booklet]
[Booklet]
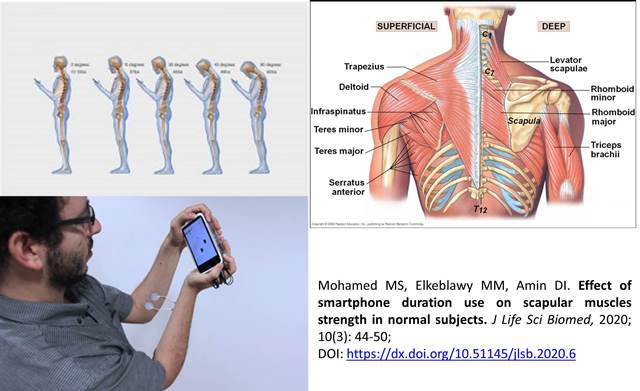
Effect of smartphone duration use on scapular muscles strength in normal subjects
| Effect of smartphone duration use on scapular muscles strength in normal subjects |
Mohamed MS, Elkeblawy MM, Amin DI.
J. Life Sci. Biomed., 10(4): 44-50, 2020; pii:S225199392000006-10
DOI: https://dx.doi.org/10.51145/jlsb.2020.6
Abstract
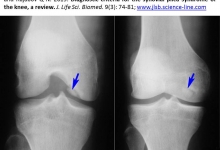
Background. The use of hand-held devices such as smartphone has been associated with shoulder pain and scapular muscles imbalance as a result of hyperactivity and tightness. Aim. Purpose of this study was to investigate the effect of smartphone duration use on pain of the upper back and scapular muscles strength in normal subjects, Methods. This study was cross sectional observational study; Eighty normal adults 20-30 years age, with right hand dominance were recruited for this study. The subjects must have at least 6 months experience in using smartphone and divided into two groups: Group A used smartphone less than 4 hours daily and group B used smartphone more than 4 hours daily. Subjects was assessed once time. Upper back pain was assessed by Visual Analogue Scale (VAS) scale and strength assessed by break test through pull and push dynamometer. Results. subjects in the two groups showed significant pain accentuation after smartphone usage, depends on the duration. Furthermore, changes in pain severity with smartphone use were different between the two groups (P<0.05). For scapular adductor muscles strength, the right dominant side was diminished but not reach to cause significant difference (P>0.05) and the left side have increase in strength with significant difference (P<0.05). Conclusion. Smartphone continuous use for more than 4 hours daily led to increase shoulder or parascapular pain and decrease strength of scapular adductor muscles in right dominant side due to prolonged hyperactivity, then led to weakness and increase the strength of left side due to hyperactivity of left side during holding or static postures and bilateral hand texting.
Keywords: Upper back pain, Smartphone, Scapular muscle strength, Pull and push dynamometer, Healthy subjects
[Full text-PDF] [HTML] [ePub] [XML] [Export citation to BibTeX, RIS & EndNote] [How to Cite] [Semantic Scholar] [Dimensions ID]
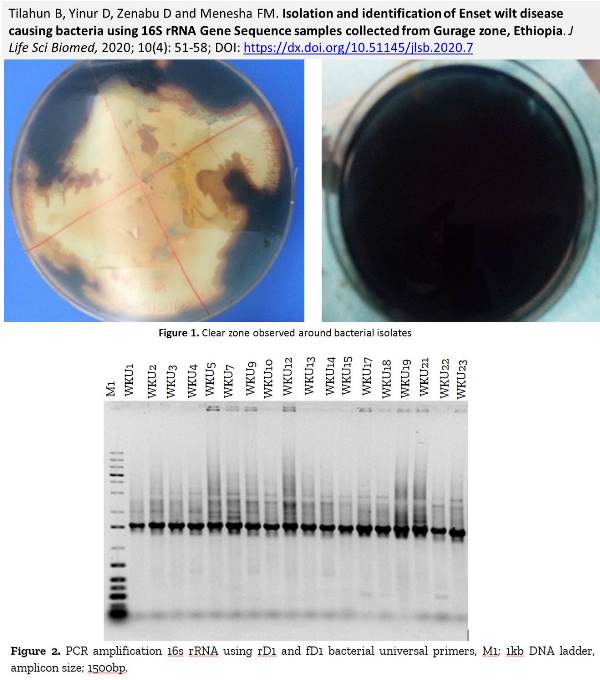
Isolation and identification of Enset wilt disease causing bacteria using 16S rRNA Gene Sequence samples collected from Gurage zone, Ethiopia
| Isolation and identification of Enset wilt disease causing bacteria using 16S rRNA Gene Sequence samples collected from Gurage zone, Ethiopia |
Tilahun B, Yinur D, Zenabu D and Menesha FM.
J. Life Sci. Biomed., 10(4): 51-58, 2020; pii:S225199392000007-10
DOI: https://dx.doi.org/10.51145/jlsb.2020.7
Abstract
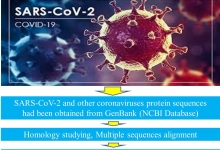
Introduction. Xanthomonas campestris is an important bacterium responsible for bacterial wilt disease, which causes predominantly a serious loss in enset production. In some enset-growing areas of Ethiopia, farmers are enforced to replace perennial enset plants with annual crops because of this disease devastates enset production. Aim. Therefore, the study aimed to identify the molecular diversity of bacterial wiealt diseas cosing bacteria from infected enset plants that were collected from the Gurage Zone, using the 16S rRNA gene sequence. Methods. 60 infected enset samples were collected from infected enset plants. Presumptive identification of the bacterium was done through biochemical tests. 16S rRNA genes of bacterial isolates were amplified using the bacteria universal primers and the amplified products were sequenced at MRC-Holland, Amsterdam. Sequence analysis and comparison were conducted to identify the isolated microbes into species and strain levels. Results. Based on the biochemical tests, 18 bacterial isolates were motile, indole negative as well as citrate and catalase positive and they were hydrolyzed starch. The sequence analysis revealed that from 18 bacterial isolates 17 of them were identified as Xanthomonas campestris of different strains and one isolate was identified as an uncultured bacterium. In this study, different Xanthomonas campestris strains that have different virulence factors were identified in the study area. To effectively control and manage bacterial wilt disease of enset plant, it is important to examine antipathogenic agent or biological control mechanisms for all Xanthomonas campestris strains. Additionally, determining plant bacterial interaction using molecular tools and identify the virulence genes are also beneficial.
Keywords: Xanthomas sp., Enset, 16srRNA gene, DNA sequencing, Wilt disease
[Full text-PDF] [HTML] [ePub] [XML] [Export citation to BibTeX, RIS & EndNote] [How to Cite] [Google Scholar] [Dimensions ID]